
(a)
Interpretation:
The structure of C from the given illustration and the explanation for how it is obtained is to be stated.
Concept introduction:
Trypsin is an enzyme that is found in the
Answer to Problem 27.53AP
The structure of C from the given illustration is
Explanation of Solution
The molecular mass of amino acid
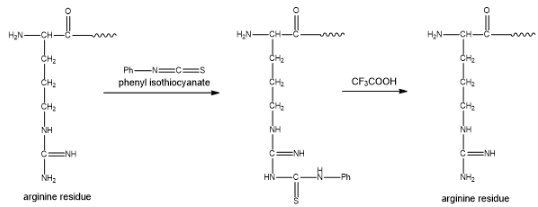
Figure 1
As a result, the phenyl hydantoin derivative is not formed. Due to this, the peptide C remain unchanged on Edman degradation. When the peptide C reacts with trypsin enzyme, it forms a single peptide D. Also, the trypsin enzyme cleaves at lysine and arginine residue. The peptide D gives the same amino acid analysis as peptide C. This confirms that the peptide D does not contain arginine residue, due to this the amino acid analysis of the peptide C is similar to that of the peptide D. The partial structure of peptide D is found to be
The structure of C from the given illustration is
(b)
Interpretation:
The b-type fragmentation for
Concept introduction:
In mass spectroscopy, compounds can be identified on the basis of the mass of the compound. When the compound breaks into fragment then they can be distinguished from the other compounds. This technique is also used to differentiate the isotopes of compounds. In amino acids, three types of fragments are observed in low energy collisions are a, b and y ions. It is known as tandem mass spectrometry.
Answer to Problem 27.53AP
The b-type fragmentation for
The b-type fragmentation for
where
Explanation of Solution
In amino acids, b-type fragments appear due to an amino group or in other words charge is being carried by N-terminal. That is why it is also known as the N-terminus amino acid fragment. The b-type fragment is shown below.
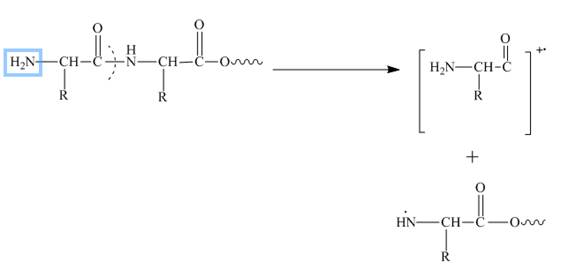
Figure 2
The peptide C is
The peptide D is
The b-type fragmentation for
Want to see more full solutions like this?
Chapter 27 Solutions
Organic Chemistry
- 22-35 Why is histidine considered a basic amino acid when the pKa of its side chain is 6.0?arrow_forward22-61 Polyglutamic acid (a polypeptide chain made only of glutamic acid residues) has an a-helix conformation below pH 6.0 and a random-coil conformation above pH 6.0. What is the reason for this conformational change?arrow_forward22-42 (a) How many atoms of the peptide bond lie in the same plane? (b) Which atoms are they?arrow_forward
- 22-47 How many different tetrapeptides can be made (a) if the peptides contain the residues of asparagine, proline, serine, and metbionine and (b) if all 20 amino acids can be used?arrow_forwardA 1.00-mg sample of a pure protein yielded on hydrolysis 0.0165 mg of leucine and 0.0248 mg of isoleucine. What is the minimum possible molar mass of the protein? (MMleucine=MMisoleucine=131g/mol)arrow_forwardUsing both three- and one-letter codes for amino acids, write the structures of all possible peptides containing the following amino acids: (a) Val, Ser, Leu (b) Ser, Leu2, Proarrow_forward
- 22-71 Which amino acid side chain is most frequently involved in denaturation by reduction?arrow_forward22-21 Explain why an amino acid cannot exist in an un-ionized form at any pH.arrow_forwardDetermine the amino acid sequence of a polypeptide from the following data:Acid-catalyzed hydrolysis gives Ala, Arg, His, 2 Lys, Leu, 2 Met, Pro, 2 Ser, Thr, and Val.Carboxypeptidase A releases Val.Edman’s reagent releases PTH-Leu.Treatment with cyanogen bromide gives three peptides with the following amino acid compositions:1. His, Lys, Met, Pro, Ser 2. Thr, Val 3. Ala, Arg, Leu, Lys, Met, Serarrow_forward
- Determine the amino acid sequence of a polypeptide from the following data: Complete hydrolysis of the peptide yields Arg, 2 Gly, Ile, 3 Leu, 2 Lys, 2 Met, 2 Phe, Pro, Ser, 2 Tyr, and Val. Treatment with Edman’s reagent releases PTH-Gly. Carboxypeptidase A releases Phe. Treatment with cyanogen bromide yields the following three peptides: 1. Gly-Leu-Tyr-Phe-Lys-Ser-Met 2. Gly-Leu-Tyr-Lys-Val-Ile-Arg-Met 3. Leu-Pro-Phe Treatment with trypsin yields the following four peptides: 1. Gly-Leu-Tyr-Phe-Lys 3. Val-Ile-Arg 2. Ser-Met-Gly-Leu-Tyr-Lys 4. Met-Leu-Pro-Phearrow_forwardDetermine the amino acid sequence of a polypeptide from the following data:Complete hydrolysis of the peptide yields Ala, Arg, Gly, 2 Lys, Met, Phe, Pro, 2 Ser, Tyr, and Val.Treatment with Edman’s reagent releases PTH-Val.Carboxypeptidase A releases Ala. Treatment with trypsin yields the following three peptides:1. Gly, Lys, Met, Tyr 2. Ala, Lys, Phe, Pro, Ser 3. Arg, Ser, Valarrow_forwardDetermine the amino acid sequence of a polypeptide from the following data:Complete hydrolysis of the peptide yields Arg, 2 Gly, Ile, 3 Leu, 2 Lys, 2 Met, 2 Phe, Pro, Ser, 2 Tyr, and Val.Treatment with Edman’s reagent releases PTH-Gly.Carboxypeptidase A releases Phe. Treatment with cyanogen bromide yields the following three peptides:1. Gly-Leu-Tyr-Phe-Lys-Ser-Met 2. Gly-Leu-Tyr-Lys-Val-Ile-Arg-Met 3. Leu-Pro-Phearrow_forward
 Chemistry: Principles and ReactionsChemistryISBN:9781305079373Author:William L. Masterton, Cecile N. HurleyPublisher:Cengage Learning
Chemistry: Principles and ReactionsChemistryISBN:9781305079373Author:William L. Masterton, Cecile N. HurleyPublisher:Cengage Learning Organic And Biological ChemistryChemistryISBN:9781305081079Author:STOKER, H. Stephen (howard Stephen)Publisher:Cengage Learning,
Organic And Biological ChemistryChemistryISBN:9781305081079Author:STOKER, H. Stephen (howard Stephen)Publisher:Cengage Learning, General, Organic, and Biological ChemistryChemistryISBN:9781285853918Author:H. Stephen StokerPublisher:Cengage Learning
General, Organic, and Biological ChemistryChemistryISBN:9781285853918Author:H. Stephen StokerPublisher:Cengage Learning Organic ChemistryChemistryISBN:9781305580350Author:William H. Brown, Brent L. Iverson, Eric Anslyn, Christopher S. FootePublisher:Cengage Learning
Organic ChemistryChemistryISBN:9781305580350Author:William H. Brown, Brent L. Iverson, Eric Anslyn, Christopher S. FootePublisher:Cengage Learning Introduction to General, Organic and BiochemistryChemistryISBN:9781285869759Author:Frederick A. Bettelheim, William H. Brown, Mary K. Campbell, Shawn O. Farrell, Omar TorresPublisher:Cengage Learning
Introduction to General, Organic and BiochemistryChemistryISBN:9781285869759Author:Frederick A. Bettelheim, William H. Brown, Mary K. Campbell, Shawn O. Farrell, Omar TorresPublisher:Cengage Learning





