
Concept explainers
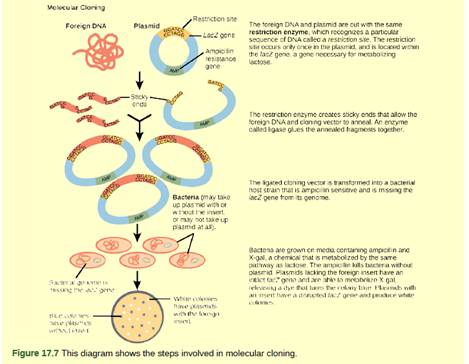
Figure 17.7 You are working in a molecular biology lab and, unbeknownst to you, your lab partner left the foreign genomic DNA that you are planning to clone on the lab bench overnight instead of storing it in the freezer. As a result, it was degraded by nucleases, but still used in the experiment. The plasmid, on the other hand, is fine. What results would you expect from your molecular cloning experiment?
- There will be no colonies on the bacterial plate.
Introduction:
Molecular cloning is a process to reproduce desired fragments of the genome. In cloning the plasmid molecules can be used to provide a folder in which desired genomic fragment is inserted. Then plasmids are introduced into a bacterial host. Plasmids with foreign DNA into them are called recombinant DNA molecules.
Answer to Problem 1VCQ
Correct answer:
The correct answer is option (b)- there will be blue colony only.
Explanation of Solution
Explanation for the correct answer:
Option (b) is given as ‘there will be blue colony only’. In this experiment, when the cloning vector plasmid with degraded foreign DNA is transformed into a bacterial host strain, releases a dye which turns the colony blue. Therefore, option (b) is correct.
Explanation for incorrect answers:
Option (a) is given as ‘there will be no colonies on the bacterial plate’. The colonies will be present on the plate. So, it is a wrong answer.
Option (c) is given as ‘there will be blue and white colonies no colonies’. Both blue and white colonies are only possible when plasmids are with foreign DNA insert and without insert.
So, it is a wrong answer.
Option (d) given as ‘there will be white colony only’. White colonies are not possible as plasmids with insert are not capable to express their characters. So, it is a wrong answer.
Hence, options (a ), (c), and (d) are incorrect.
The molecular cloning is a process to reproduce the desired fragments of the genome is reproduced by molecular cloning with the help of plasmid which provides a folder in which desired genomic fragment is inserted. Hence, the correct answer is option (b).
Want to see more full solutions like this?
Chapter 17 Solutions
Biology 2e
Additional Science Textbook Solutions
College Physics
Biological Science
Human Physiology: An Integrated Approach (8th Edition)
Anatomy & Physiology (6th Edition)
Concepts of Genetics (11th Edition)
Essentials of Genetics (9th Edition) - Standalone book
- Scientists modified the tumor-inducing (Ti) plasmid, a naturally occurring plant plasmid found in Agrobacterium tumefaciens, to create a tool used to introduce any gene of interest into plant cells. They created a binary system because a single Ti plasmid is too large to be easily manipulated. One part of the system is a disarmed plasmid, and the second part is a transformation vector. The following sentences describe the function of key DNA elements in the system. Virulence region Genes for conjugative transfer Disarmed Ti plasmid (T-region removed) ori Kan selectable marker plant selectable marker constitutive promoter T-region conjugative transfer virulence Kan (bacterial selectable marker) Amp selectable marker Plant selectable marker (e.g., herbicide resistance) Constitutive promoter T-DNA left border Place the terms in the appropriate blanks to complete the sentences. Not all terms will be used. 3' transcriptional terminator MCS (inserted gene of interest) T-region Transformation…arrow_forwardA molecular geneticist hopes to find a gene in human liver cells that codes for an important blood-clotting protein. He knows that the nucleotide sequence of a small part of the gene is CTCGACTCACA. Briefly explain how to obtain the desired gene. Briefly describe how to clone the desired gene into a cloning plasmid.arrow_forwardYou are asked to sequence a piece of DNA to determine if it is from a gene thought to be involved in the development of breast cancer. The sequence of the template strand is ATGCCCGTAATCGTTA and you are given the primer TAACGA. You take these along with a sequencing kit that contains everything else needed for sequencing. You then run the sequencing experiment and analyze the results on a sequencing gel. Which of the following gels (A-D) is the correct sequencing gel for this experiment? Answers: A-D A A BB CC DD Question #3 attachment A ATOC B с A TO C ATOC ATOCarrow_forward
- After transformation you were asked to grow bacterial cells transformed with plasmid on a plate that had X-gal and ampicillin. X-Gal is often used as in indicator dye, which turns blue when metabolized by B-galactosidase protein and used to test if cloning experiments have worked. [Note look at the vector diagrams carefully] Briefly explain how you would find the bacterial cells that are transformed with the plasmid with the YFG inserted.arrow_forwardIf I clone a complete eukaryotic gene, including the eukaryotic promoter region, ligate it into a plasmid, and transform it into E. coli, will I be able use the transformed E. coli to make the corresponding protein? Explain why, or why not? If you decide to do this, what would your cloning strategy be?arrow_forwarda)What two restriction sites are you going to use to clone your PCR product into the pL4440 plasmid? What are their DNA sequence? b) State the primer sequence you will use to amplify the F27C1.7 gene ready to be cloned into the pL4440 plasmid? c) How would you go about cloning this amplified DNA into pL4440? Using your knowledge of cloning list 5 important aspects of the method.arrow_forward
- You want to clone a 6,000 bp DNA fragment in E. coli. Which cloning vectors would be appropriate? How will you select transformants?arrow_forwardIn the following "gene library" cloning experiment. Digested genomic DNA AmpR gene TCR gene TCR is tetracycline resistant marker, AmpR is ampicillin resistant marker and BamHI is the unique restriction enzyme on plasmid. Cells transformed with the "recombinant plasmids" will be:....... (Hint: this is a challenging question) a) resistant to ampicillin and tetracycline b) sensitive to tetracycline and ampicillin c) resistant to tetracycline and sensitive to ampicillin d) resistant to ampicillin and sensitive to tetracycline e) sensitive to ampicillin and tetracycline BamHIarrow_forwardYou’re working in a research lab, and your current task is to clone the gene that codes for tyrosinase from potatoes. You grind up some potato, extracts the DNA from it and digests the DNA with two different restriction enzymes (separately, not together): EcoRI and BamHI. You then obtain the cloning vector, pUC19, and digest it with the same two enzymes. You then run a gel which is shown here. You notice that the cloning vector made nice, tight bands on the gel, but the potato DNA just looks like a smear with no distinct bands. However, this is just what you expected. Explain why there are so many bands. Which enzyme would be the better choice to use for cloning the potato DNA, EcoRI, or BamHI? Explain why? Be specific.arrow_forward
- With the use of well-illustrated diagrams, reconstruct the entire cloning process by explaining different stages of the cloning process that involves the following:a. Isolation of target DNA fragments (often referred to as inserts)b. Ligation of inserts into the plasmid, creating recombinant molecules c. Transformation of recombinant plasmids into bacteria or other suitable host for propagationd. Screening/selection of hosts containing the intended recombinant plasmid. For this stage(d), discuss the importance of a second marker that can be used for screening of genomic DNA for colonies containing the pka-1 under the principle of insertional inactivation. This should be properly explained using all the attributes of the plasmid described above.arrow_forwardYou are studying a new plasmid, and you digest the plasmid with three restriction enzymes: Eco RI (E), HindlII (H), and Xbal (X). You digest the plasmid DNA with each of the following combinations of enzymes and observe the results on an agarose gel. You are provided a partial plasmid map as shown below to the right. E+H E+X H+x Kb +4.3 +2.8 -+2.5 -2.0 - -1.8 +1.5 -1.0 12 F0.8 +0.5 a. What is the size of this plasmid in base pairs? b. What is the distance in base pairs between E1 and H? c. What is the distance in base pairs between E1 and X? d. What is the distance in base pairs between E2 and H? e. What is the distance in base pairs between E2 and X?arrow_forwardFigure 2 illustrates the important elements of a cosmid.a) Briefly describe the importance of origin of replication in a cosmid and where are thecos sites derived from.b) The cloning capacity of a cosmid is up to 44 kilo base pairs. State TWO classes of DNAcloning vectors that have higher cloning capacity.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
